Introduction
Pneumonia is an important socio-medical problem and one of the leading causes of death in the world1. Gram-negative rods, crucial pathogens in hospital- as well as community-acquired pneumonia, produce and release endotoxins which constitute lipopolysaccharides (LPS). Once inhaled via respiratory routes, LPS stimulate alveolar structural as well resident cells to release many kinds of bioactive agents such as proinflammatory cytokines into local microenvironments2.
Recent studies have emphasized a crucial role for the lung epithelium as an important sentinel and effector system of innate immunity3–5. Upon infectious agents and their products being inhaled into the respiratory systems, activated lung epithelium may contribute to the regulation of the immune response as the first-line defense mechanism6. Among those defense responses, it seems important that alveolar epithelial cells express and release a variety of pro-inflammatory cytokines and chemokines into alveolar microenvironments6,7. A CXC chemokine, interleukin (IL)-8, plays an important role in the acute recruitment of immune/inflammatory cells, especially neutrophils, to the site of infection in the lung8,9. LPS bind to Toll-like receptors (TLR) and thereby activate downstream signal transduction pathways which ultimately phosphorylate cytosolic I-κB kinase10,11. Then, I-κB is phosphorylated to induce free-form of NF-κB, which translocates into the nucleus. The NF-κB binds to its specific binding sites on the promoter regions and enhances the expression of IL-8 gene12,13.
Increasing evidence has indicated that the expression of many inflammatory genes involves the remodeling of the chromatin structure provided by histone proteins14,15. Remodeling of chromatin within the nucleus is controlled by the degree of acetylation/deacetylation of histone residues on the histone core around which DNA is coiled. Histone acetylation results in the unwinding of the chromatin structure, which enhances the binding of transcription factors to their specific promoter sites on the DNA16. Nuclear histone acetylation is a reversible process and is regulated by a group of histone acetyltransferases (HATs) which promote acetylation, and histone deacetylases (HDACs) which promote deacetylation17,18. The loosening of DNA-histone interactions and the subsequent unmasking of transcription factor binding sites is controlled by specific covalent modifications of accessible N-terminal histone tails19. Among the four core histone proteins that comprise the central chromatin core (H2A, H2B, H3, and H4), acetylation processes on H3 and H4 seem particularly important in gene regulation. For example, Gilmour and associates20 found that acetylation on H4 played an important role in environment particle-induced IL-8 production in A549 cells. Viable Listeria monocytogenes-stimulated endothelial cells showed increased expression of IL-8, and that process depended on modifications of H3 and H421. Although the host response in pneumonia is characterized by massive cytokine production, and altered histone modifications have been observed in diseased lungs22, it is not fully elucidated how histone modifications contribute to innate immune regulation in the lung.
In this study, we tried to determine whether Escherichia coli-derived LPS, one of the mainstream stimuli upon bacterial respiratory infection, altered histone acetylation/deacetylation balance, and to see whether the modulation of HDACs or HATs by their specific inhibitors (i.e. trichostatin A [TSA] for HDACs and anacardic acid for HATs) affected IL-8 gene expression and protein production in an alveolar epithelial cell line A549 in vitro.
Materials and methods
Cell culture and stimulation
Human alveolar epithelial cell line A549 was obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA) via DS Pharma Biomedical Co., Ltd (Tokyo, Japan). A549 cells were cultured in Dulbecco’s Modified Eagle’s Medium (Sigma-Aldrich, St. Louis, MO, USA) containing 10% fetal bovine serum (FBS) (Invitrogen, Grand Island, NY, USA) and 1% penicillin-streptomycin (Sigma-Aldrich, St, Louis, MO, USA), and incubated at 37°C in 5% CO2 atmosphere. Cells were cultured to 80% confluency as judged under inverted microscopy before the medium was replaced with serum-free Dulbecco’s Modified Eagle’s Medium and incubated for a further 15 h. The cells were stimulated with different concentrations of E.coli-derived LPS (026:B6, Sigma-Aldrich, St, Louis, MO, USA) for further experiments. To evaluate the effects of HDAC and HAT inhibitors, cells were pre-treated with TSA or anacardic acid (Sigma-Aldrich, St, Louis, MO, USA; dissolved in dimethyl sulfoxide [DMSO] and further diluted for use) at the concentrations indicated, 1h prior to stimulation with LPS (10 μg/ml).
Quantitative reverse transcription polymerase chain reaction (qRT-PCR)
Total RNA from A549 cells was isolated with the RNeasy Mini kit (Qiagen, Hamburg, Germany), and cDNA was prepared using the Im Prom II reverse transcription system (Promega, Madison, WI, USA) for reverse transcription, and all procedures were conducted according to the manufacturers’ instructions. Gene transcript levels of IL-8, and glyceraldehyde-3-phosphate dehydrogenase (GAPDH) were quantified by real-time PCR using a reaction mixture with SYBR Premix Ex Taq (Takara, Tokyo, Japan) on a 7500 real-time PCR system (Applied Biosystems, Foster City, CA, USA). The primer sets for IL-8 were: (forward) 5’-CTGATTTCTGCAGCTCTGTG-3’; (reverse) 5’-TTCACTGGCATCTTCACTG-3’ and for GAPDH were: (forward) 5’-TGAACGGGAAGCTCACTGG-3’ (reverse) 5’-TCCACCACCCTGTTGCTGTA-3’ The relative amount of gene transcript was estimated after normalization by dividing the calculated value for the gene of interest by the GAPDH value.
Chromatin immunoprecipitation assay (ChIP assay)
Chromatin immunoprecipitation was performed using Millipore’s ChIP kit (Billerica, Massachusetts, USA) with an acetyl-histone H4 antibody. Cells were cultured to 80% confluency in 100mm culture plates. Cells were cross-linked by adding 1% formaldehyde for 10 minutes at room temperature in shaking. Then, 1ml of 10×glycine was added to 10ml of growth media to each dish to quench unreacted formaldehyde at room temperature for 5 minutes. Cells were washed twice with cold 10ml phosphate buffered saline (PBS). One ml PBS containing 1×protease inhibitor cocktail II prepared in the ChIP kit was added to each dish. Cells were scraped from each dish into a conical tube. The tubes were centrifuged at 700×g at 4°C for 5 minutes to pellet cells. Each cell pellet was resuspended in 1ml of SDS lysis buffer containing 1×protease inhibitor cocktail II prepared in the ChIP kit. Chromatin was sonicated to an average DNA length of 200–1000 bp using an Ultrasonic Disruptor UD-200 sonicator (TOMY, Tokyo, Japan). Sonicated samples were centrifuged and the supernatant was collected. Samples (100μl) of the extracted chromatin were diluted with 900μl ChIP Dilution Buffer (0.01% SDS, 1.1% Triton X-100, 1.2mM EDTA, 16.7mM Tris-HCl, pH 8.1, 167mM NaCl), precleared (1 hour) by incubation with 60μl Protein G Agarose containing 1×protease inhibitor cocktail II, and subjected to immunoprecipitation with a specific antibody with rotation overnight at 4°C. The antibody used for ChIP assays was anti-acetyl histone H4 antibody (Millipore, Billerica, Massachusetts, USA). Immunocomplexes were collected by adsorption onto 60μl Protein G Agarose and the beads were washed five times sequentially with Low Salt Immune Complex Wash Buffer (0.1% SDS, 1% Triton X-100, 2mM EDTA, 20mM Tris-HCl, pH 8.1, 150mM NaCl), High Salt Immune Complex Wash Buffer (0.1% SDS, 1% Triton X-100, 2mM EDTA, 20mM Tris-HCl, pH 8.1, 500mM NaCl) and LiCl Immune Complex Wash Buffer (0.25M LiCl, 1% IGEPAL CA630, 1% deoxycholic acid (sodium salt), 1mM EDTA, 10mM Tris, pH 8.1), which were prepared in the ChIP kit. Precipitates were washed twice with TE Buffer (10mM Tris-HCl, pH 8.0, 1mM EDTA), and antibody-chromatin fragments were eluted from the beads with 1% sodium dodecyl sulphate in 0.1 M NaHCO3. Cross-links were reverted by adding 200mM NaCl and heating at 65°C for 5 hours. In total, 10mg/ml RNase A were added and samples were then incubated for 30 minutes at 37°C. 10mg/ml proteinase K, 10mM EDTA and 40mM Tris-HCl were added and samples were then incubated for 2 hours at 45°C.
A total of 1ml of Bind Reagent A was added to each sample, and mixed well by pipetting. The sample was transferred up to a spin filter placed in a collection tube, and was centrifuged for 30 seconds at 12000×g. Then, 500μl of Wash Reagent B was added to each spin filter placed in a collection tube, and was centrifuged for 30 seconds at 12000×g. Then, 50μl of Elution Buffer C was added to each spin filter placed in a clean collection tube, and was centrifuged for 30 seconds at 12000×g to recover purified DNA. The IL-8 enhancer regions were quantified by real-time PCR using a reaction mixture with SYBR Premix Ex Taq (Takara, Tokyo, Japan) on a 7500 real-time PCR system (Applied Biosystems, Foster City, CA, USA). The primer sets for IL-8 were: (forward) 5’- CAGAGACAGCAGAGCACAC-3’; (reverse) 5’- ACGGCCAGCTTGGAAGTC-3’. All PCR signals from immunoprecipitated DNA were normalized to PCR signals from non-immunoprecipitated input DNA.
ELISA for IL-8 measurement
The concentration of IL-8 in the media was determined by sandwich ELISA kit (R&D Systems, San Diego, CA) according to the manufacturer’s instructions.
Statistical analysis
Data were analyzed with the Statistical Package for Social Science (SPSS) version 17.0 for Windows (SPSS Inc., Chicago, IL, USA). Values are expressed as mean ± standard error (SE). Mann-Whitney U test was performed for comparisons between groups. A P-value < 0.05 was considered significant. All P-values were two-sided.
Results
LPS stimulated production of IL-8 in A549 cells
We analyzed IL-8 production in LPS stimulated A549 cells. A549 cells were stimulated by different concentrations of LPS (10 ~ 200μg/ml) for indicated time periods. As shown in Figure 1, LPS showed a time- and dose-dependent stimulatory effect on IL-8 release.
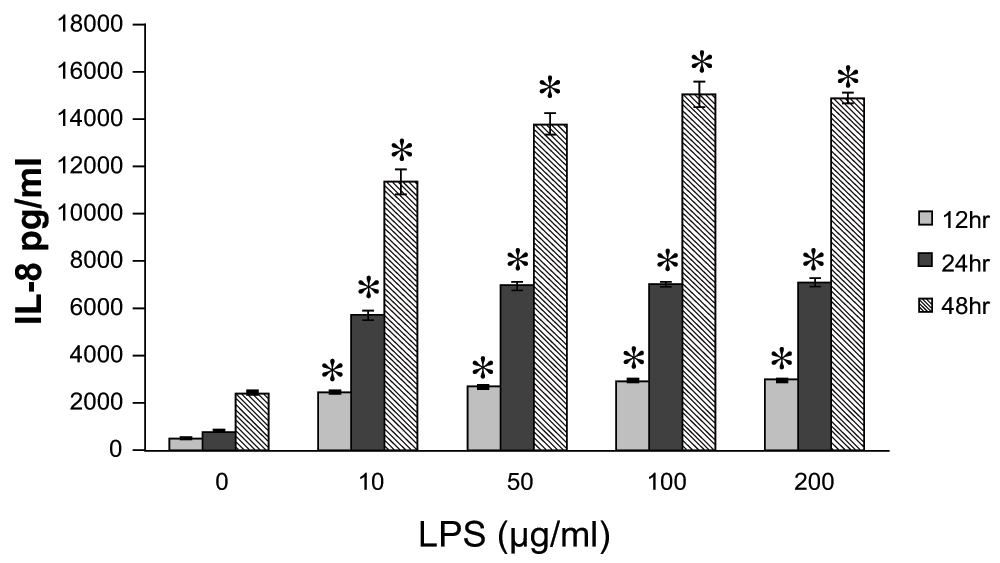
Figure 1. The dose- and time- dependent interleukin-8 (IL-8) release from A549 cells in vitro.
A549 cells were cultured until 80% confluence and stimulated by different concentrations of lipopolysaccharide (LPS) (10 ~ 200μg/ml). LPS showed a time- and dose-dependent stimulatory effect on IL-8 release at each concentration. Data were expressed in mean±SEM. *P<0.05, compared to LPS unstimulated cells at each concentration and time point (Mann-Whitney U test), n=4.
LPS induced expression of IL-8 gene in A549 cells
A549 cells were stimulated by 10μg/ml LPS, and the time course in levels of IL-8 mRNA was analyzed by qRT-PCR. IL-8 mRNA levels showed a gradual increase in response to LPS, reaching a maximum level 2 h after initial stimulation with 10μg/ml, which then decreased after that point (Figure 2).
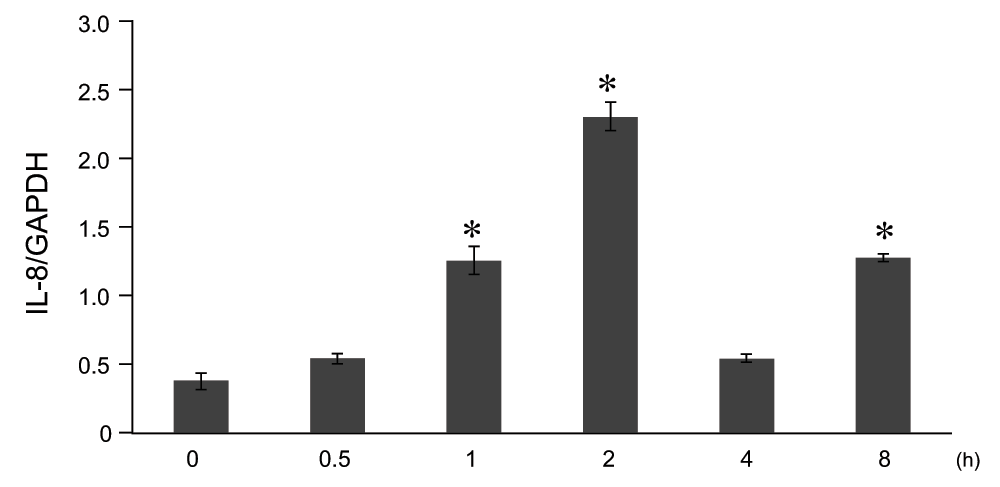
Figure 2. Time course analysis of lipopolysaccharide (LPS)-mediated interleukin-8 (IL-8) gene expression in A549 cells in vitro.
A549 cells were cultured until subconfluence and stimulated by LPS at 10μg/ml. After 0 ~ 8 hrs, IL-8 mRNA expression was analyzed by quantitative reverse transcription polymerase chain reaction (qRT-PCR). The IL-8 mRNA levels were normalized to glyceraldehyde-3-phosphate dehydrogenase (GAPDH) levels. Data were expressed in mean±SEM. *P<0.05: compared to LPS 0hr stimulated cells (Mann-Whitney U test), n=4.
LPS induced histone acetylation in A549 cells
Based on the previous reports showing that modifications of histones, in particular acetylation of H4, seem to contribute to the regulation of inflammatory genes such as IL-820, we assessed acetylation of H4 after stimulation of LPS in A549 cells. We analyzed histone modifications at the IL-8 gene promoter by ChIP assay. A549 cells were stimulated with LPS (10μg/ml) for 0 min to 3 h. LPS induced a time-dependent increase of acetylation of H4 at the IL-8 promoter, and this increase peaked after 1 h (P<0.05, Mann-Whitney U test), and then decreased after 3 h of LPS stimulation (Figure 3).
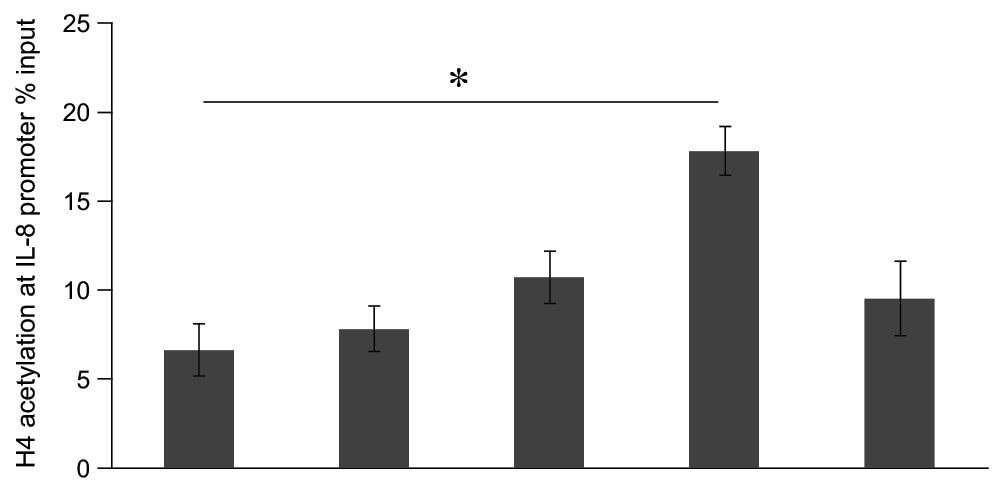
Figure 3. Time course analysis of lipopolysaccharide (LPS)-induced histone H4 acetylation at the promoter region of the interleukin-8 (IL-8) gene in A549 cells in vitro.
A549 cells were stimulated 0 ~ 3 hr with LPS (10μg/ml). The results presented are from ChIP analyses using anti-acetyl H4 antibodies. All PCR signals from immunoprecipitated DNA were normalized to PCR signals from non-immunoprecipitated input DNA. Results are expressed as percentage of the input. Data were expressed in mean±SEM. *P<0.05: compared to LPS 0hr stimulated cells (Mann-Whitney U test), n=4.
Histone acetylation regulates IL-8 expression in LPS-stimulated A549 cells
Next, we wondered whether inhibition of HDACs by TSA or blocking of HATs by anacardic acid impacts on LPS-induced IL-8 expression. We increased global histone acetylation by incubation of A549 cells with TSA (10nM, 24 h) which did not induce IL-8 secretion per se (Figure 4). Preincubation for 1 h with TSA (10nM) before subsequent treatment with LPS significantly increased IL-8 release as compared to LPS alone (Figure 4). Pretreatment with TSA (10nM) showed a tendency to increase IL-8 mRNA levels as assessed by qPCR analysis, but did not reach statistical significance (Figure 5).
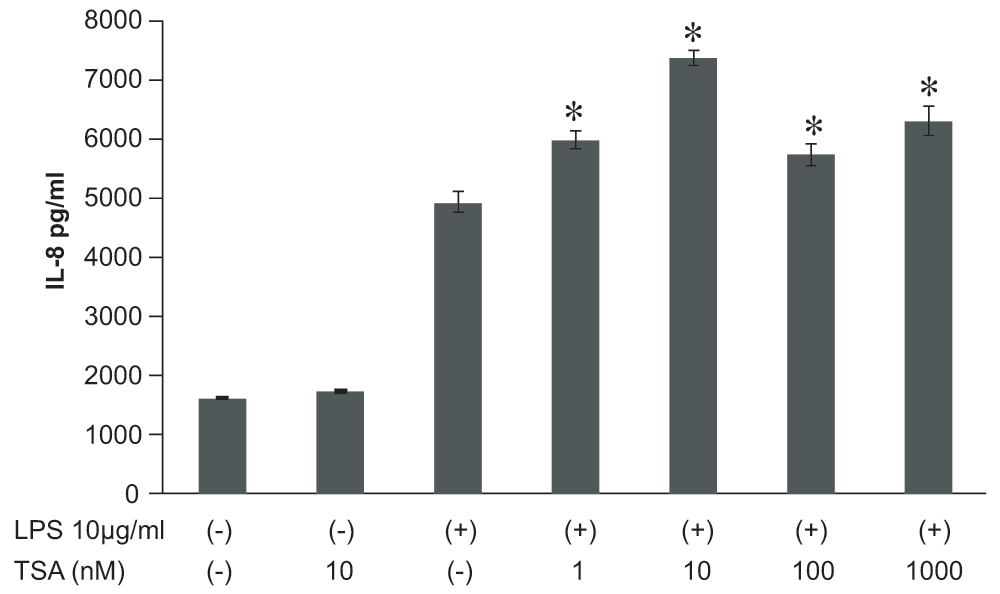
Figure 4. Effect of trichostatin A (TSA) on lipopolysaccharide (LPS)-stimulated interleukin-8 (IL-8) release from A549 cells in vitro.
TSA at 1 ~ 1000nM treated 1 hr before LPS stimulation (10μg/ml) showed a significant stimulatory effect on LPS-induced IL-8 release. Data were expressed as mean±SEM. *P<0.05: compared to LPS-stimulated cells (Mann-Whitney U test), n=4.
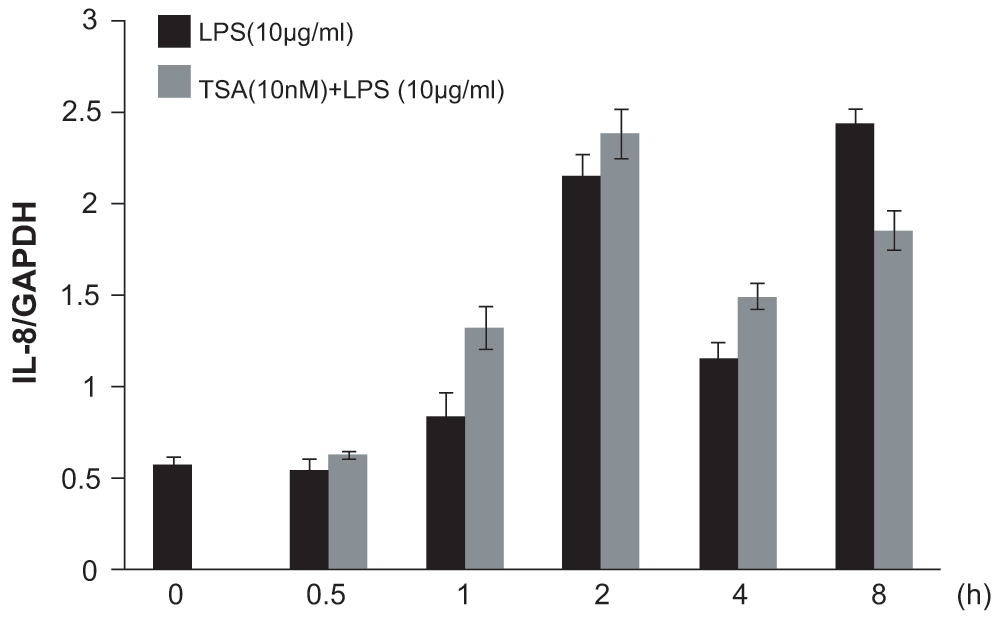
Figure 5. Time course analysis of trichostatin A (TSA) on lipopolysaccharide (LPS)-stimulated interleukin-8 (IL-8) gene activation from A549 cells in vitro.
TSA at 10nM treated 1 h before LPS stimulation (10μg/ml) tended to show a stimulatory effect on LPS-induced IL-8 gene activation. After 0 ~ 8 h, the levels of IL-8 mRNA were analyzed by quantitative reverse transcription polymerase chain reaction (qRT-PCR). The IL-8 mRNA levels were normalized to glyceraldehyde-3-phosphate dehydrogenase (GAPDH) levels, n=4.
Suppression of histone acetylation by blocking of HATs via anacardic acid showed a dose-dependent inhibitory effect on LPS-stimulated IL-8 production (Figure 6), indicating that histone deacetylation regulates IL-8 expression in LPS-treated epithelial cells. The effects of anacardic acid on IL-8 mRNA expression were analyzed by qRT-PCR. Anacardic acid at 100μM administered 1 h before LPS (10μg/ml) stimulation showed a significant inhibitory effect on LPS-induced IL-8 mRNA levels (Figure 7). These observations indicated that modulation of histone acetylation by anacardic acid regulated LPS-stimulated IL-8 gene expression in A549 cells.
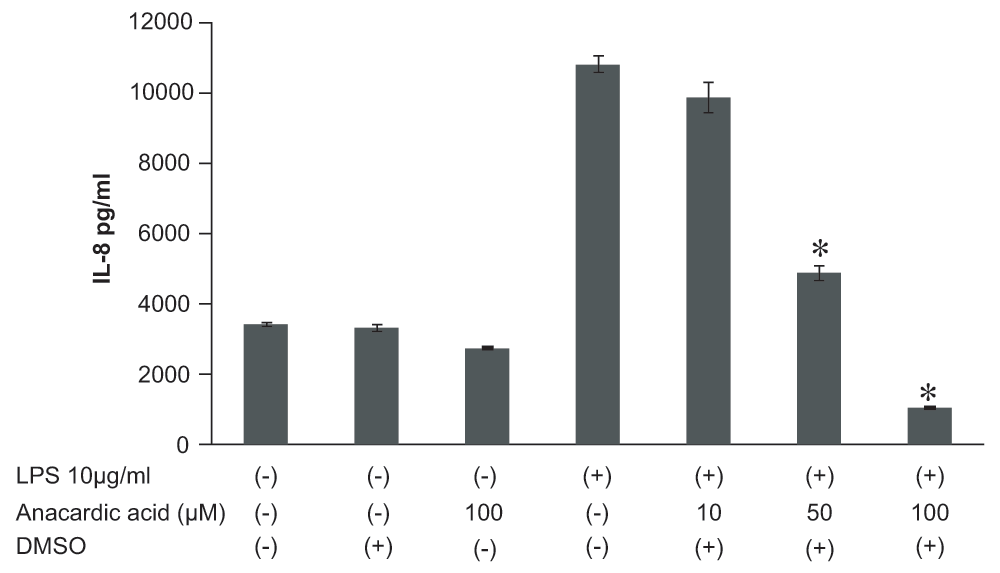
Figure 6. Effect of anacardic acid on lipopolysaccharide (LPS)-stimulated interleukin-8 (IL-8) release from A549 cells in vitro.
Anacardic acid at 10 ~ 100μM treated 1 hr before LPS stimulation showed a significant inhibitory effect on LPS-induced IL-8 release. Data were expressed as mean±SEM. DMSO: dimethyl sulfoxide used for solvent. *P<0.05: compared to LPS (10μg/ml) stimulated cells (Mann-Whitney U test), n=4.
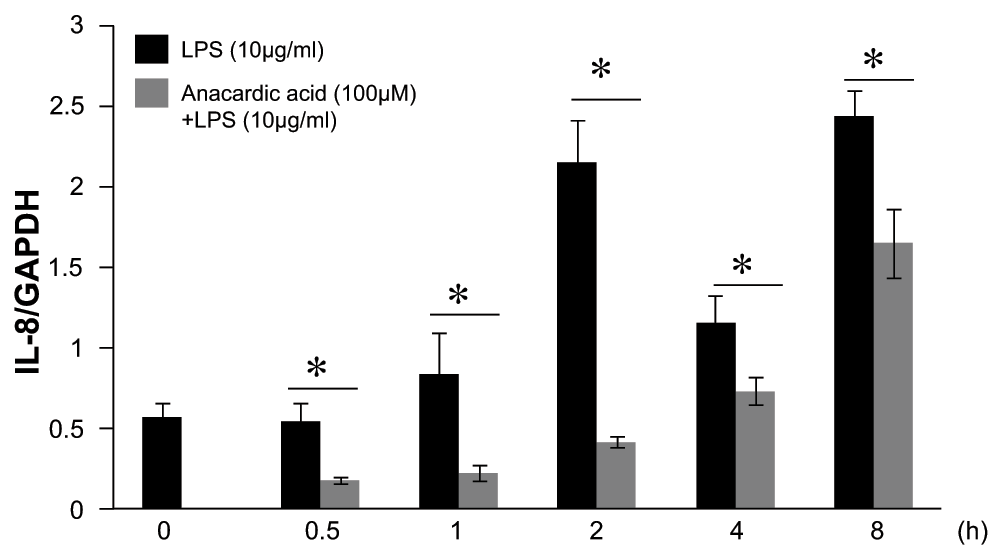
Figure 7. Time course analysis of anacardic acid on lipopolysaccharide (LPS)-stimulated interleukin-8 (IL-8) gene activation in A549 cells in vitro.
Anacardic acid at 100μM treated 1 hr before LPS (10μg/ml) stimulation showed a significant inhibitory effect on LPS-induced IL-8 gene activation. After 0 ~ 8hrs, the levels of IL-8 mRNA were analyzed by quantitative reverse transcription polymerase chain reaction (qRT-PCR). The IL-8 mRNA levels were normalized to glyceraldehyde-3-phosphate dehydrogenase (GAPDH) levels. Data were expressed as mean±SEM. *P<0.05 (Mann-Whitney U test), n=4.
Discussion
In the present study, we demonstrated that E. coli-derived LPS stimulate human alveolar epithelial A549 cells to express and release IL-8, an important chemokine for the local recruitment of neutrophils as an initial defense mechanism8,9. The specific histone acetylation processes were evaluated by ChIP assay, which clearly showed that LPS induced histone H4 acetylation in A549 cells. Next, we studied the effects of TSA, a HDAC inhibitor on LPS-induced IL-8 expression. TSA showed a significant stimulatory effect on IL-8 production, whereas this agent tended to increase, although not significantly, IL-8 mRNA levels in LPS-stimulated A549 cells. Finally, a HAT inhibitor, anacardic acid, significantly decreased IL-8 mRNA levels as well as protein release in a dose-dependent fashion. These results suggested that LPS-induced IL-8 gene expression is, at least in part, regulated by histone H4 acetylation/deacetylation balance at the IL-8 promoter region.
It has been reported that alveolar epithelial cells respond to bacterial products such as LPS to produce a variety of inflammatory cytokines and mediators including IL-82. We23,24 and others9,25,26 have previously shown that airway and alveolar epithelial cells, including A549 cells, are activated by a variety of endogenous agents such as cytokines as well as exogenous stimuli including LPS and fine particles, and express biologically active compounds including cytokines and chemokines.
The transcription of many genes is known to correlate with levels of acetylated nuclear histone proteins14–16. The expression of many inflammatory genes involves the remodeling of the chromatin structure provided by histone proteins. Histone acetylation causes the unwinding of the chromatin structure, thereby enabling transcription factors to bind to their specific promoter sites on the DNA. Such acetylation processes are reversible and regulated by HATs, which promote acetylation, and HDACs, which promote deacetylation18,19.
Proinflammatory gene transcription regulation is a multifaceted process that requires integrated sequential molecular events for maximal gene transcription to occur. Activation of specific gene expression needs to be associated with co-operated chromatin remodeling by histone acetylation and binding of transcription factors to their specific binding sites on DNA. In the present experiments, the levels of histone H4 acetylation peaked 1 h after LPS stimulation followed by the maximal levels of IL-8 mRNA at 2 h; such a time course was consistent with the above scenarios.
Our study supports a hypothesis that histone H4 acetylation could play a key role in inflammatory gene transcription such as IL-8 in A549 cells caused by LPS; however, the role of acetylation of the other histones in this model is yet to be established. It has been reported that a variety of histone acetylations are involved in cytokine transcriptional processes. Miyata et al.27 have shown that H3 and H4 are acetylated in murine epithelial cells in response to granulocyte-colony stimulating factor (G-CSF) at the myeloperoxidase gene promoter site, a response dependent on MAPK activation.
Anacardic acid is a bioactive phytochemical found in the nutshell of Anacardium occidentale28. The promising effect of anacardic acid on IL-8 gene regulation in the current study has already been reported by Schmeck and associates29 in Legionella pneumophila-derived flagellin-induced IL-8 expression. Although the available records are promising, more detailed investigation into the therapeutic properties of anacardic acid, particularly the anti-cancer and anti-inflammatory activities, are needed30.
LPS-mediated IL-8 production is generally expected to act as a defense mechanism which recruits neutrophils in the local environment and facilitates bacterial elimination. However, it is now also known that excessive accumulation of activated neutrophils in the lung is likely to cause excessive inflammatory changes that damage lung tissues8. In this context, appropriate control of local inflammation would be an important strategy for the management of severe pneumonia22. Recently, it has been attracting attention that macrolide antibiotics added to guideline-based therapy has shown a better outcome than respiratory quinolones among intensive care unit (ICU) hospitalized patients with severe pneumonia and/or sepsis31. It has been speculated that certain anti-inflammatory actions of macrolides were involved in such clinically beneficial effects32. Therefore, the suppressive effects of anacardic acid on IL-8 gene expression and production might become a novel strategy for controlling excessive inflammation in order to lessen the risks of acute lung injury and, ultimately, respiratory failure to death.
In conclusion, we have shown that E. coli-derived LPS-induced expression and release of IL-8 are associated with increased acetylation of histone H4 expression. The HDAC inhibitor TSA increased IL-8 release. In contrast, a HAT inhibitor, anacardic acid, significantly decreased IL-8 mRNA levels as well as protein release in a dose-dependent fashion. These results suggested that regulation of HDAC and HAT activity by low molecular weight agents might become a novel strategy for the appropriate control of inflammation by modulating inflammatory mediators such as IL-8.
Author contributions
TY, HW, HT and HG conceived the study. TY and MH designed the experiments. TY, MN, KH, MW and HI carried out the research. SK contributed to the design of experiments and provided expertise in ChIP experiments. TY, HW and HT prepared the first draft of the manuscript. MW and HI contributed to the experimental design and preparation of the manuscript. All authors were involved in the revision of the draft manuscript and have agreed to the final content.
Competing interests
No relevant competing interests were disclosed.
Grant information
This work was supported in part by the scientific grants from the Japanese Ministry of Education, Science and Culture (Hajime Goto, M.D., Ph.D.: grant number 21591298, and Hiroo Wada, M.D., Ph.D.: grant number 23591481).
Acknowledgements
We thank Ms Makiko Sakamoto for technical assistance during this project. We also thank Ms Yumiko Date for secretarial assistance.
Faculty Opinions recommendedReferences
- 1.
Mizgerd JP:
Acute lower respiratory tract infection.
N Engl J Med.
2008; 358(7): 716–727. PubMed Abstract
| Publisher Full Text
| Free Full Text
- 2.
Thorley AJ, Grandolfo D, Lim E, et al.:
Innate immune responses to bacterial ligands in the peripheral human lung--role of alveolar epithelial TLR expression and signalling.
PLoS One.
2011; 6(7): e21827. PubMed Abstract
| Publisher Full Text
| Free Full Text
- 3.
Gilmour PS, Rahman I, Hayashi S, et al.:
Adenoviral E1A primes alveolar epithelial cells to PM(10)-induced transcription of interleukin-8.
Am J Physiol Lung Cell Mol Physiol.
2001; 281(3): L598–L606. PubMed Abstract
- 4.
Hashimoto S, Gon Y, Takeshita I, et al.:
Diesel exhaust particles activate p38 MAP kinase to produce interleukin 8 and RANTES by human bronchial epithelial cells and N-acetylcysteine attenuates p38 MAP kinase activation.
Am J Respir Crit Care Med.
2000; 161(1): 280–5. PubMed Abstract
| Publisher Full Text
- 5.
Nakamura H, Yoshimura K, Jaffe HA, et al.:
Interleukin-8 gene expression in human bronchial epithelial cells.
J Biol Chem.
1991; 266(29): 19611–7. PubMed Abstract
- 6.
Chuquimia OD, Petursdottir DH, Rahman MJ, et al.:
The Role of Alveolar Epithelial Cells in Initiating and Shaping Pulmonary Immune Responses: Communication between Innate and Adaptive Immune Systems.
PLoS One.
2012; 7(2): e32125. PubMed Abstract
| Publisher Full Text
| Free Full Text
- 7.
Sanders CJ, Doherty PC, Thomas PG:
Respiratory epithelial cells in innate immunity to influenza virus infection.
Cell Tissue Res.
2011; 343(1): 13–21. PubMed Abstract
| Publisher Full Text
- 8.
Chollet-Martin S, Montravers P, Gibert C, et al.:
High levels of interleukin-8 in the blood and alveolar spaces of patients with pneumonia and adult respiratory distress syndrome.
Infect Immun.
1993; 61(11): 4553–9. PubMed Abstract
| Free Full Text
- 9.
Li B, Dong C, Wang G, et al.:
Pulmonary epithelial CCR3 promotes LPS-induced lung inflammation by mediating release of IL-8.
J Cell Physiol.
2011; 226(9): 2398–405. PubMed Abstract
| Publisher Full Text
- 10.
Akira S:
Toll-like receptor signaling.
J Biol Chem.
2003; 278(40): 38105–38108. PubMed Abstract
| Publisher Full Text
- 11.
Mitchell JA, Paul-Clark MJ, Clarke GW, et al.:
Critical role of toll-like receptors and nucleotide oligomerisation domain in the regulation of health and disease.
J Endocrinol.
2007; 193(3): 323–30. PubMed Abstract
| Publisher Full Text
- 12.
Henkel T, Machleidt T, Alkalay I, et al.:
Rapid proteolysis of I kappa B-alpha is necessary for activation of transcription factor NF-kappa B.
Nature.
1993; 365(6442): 182–5. PubMed Abstract
| Publisher Full Text
- 13.
Park GY, Christman JW:
Nuclear factor kappa B is a promising therapeutic target in inflammatory lung disease.
Curr Drug Targets.
2006; 7(6): 661–8. PubMed Abstract
| Publisher Full Text
- 14.
Egger G, Liang G, Aparicio A, et al.:
Epigenetics in human disease and prospects for epigenetic therapy.
Nature.
2004; 429(6990): 457–63. PubMed Abstract
| Publisher Full Text
- 15.
Barnes PJ, Adcock IM, Ito K:
Histone acetylation and deacetylation: importance in inflammatory lung diseases.
Eur Respir J.
2005; 25(3): 552–63. PubMed Abstract
| Publisher Full Text
- 16.
Kuo MH, Allis CD:
Roles of histone acetyltransferases and deacetylases in gene regulation.
Bioessays.
1998; 20(8): 615–626. PubMed Abstract
| Publisher Full Text
- 17.
Tang YA, Wen WL, Chang JW, et al.:
A novel histone deacetylase inhibitor exhibits antitumor activity via apoptosis induction, F-actin disruption and gene acetylation in lung cancer.
PLoS One.
2010; 5(9): e12417. PubMed Abstract
| Publisher Full Text
| Free Full Text
- 18.
Claus R, Fliegauf M, Stock M, et al.:
Inhibitors of DNA methylation and histone deacetylation independently relieve AML1/ETO-mediated lysozyme repression.
J Leukoc Biol.
2006; 80(6): 1462–72. PubMed Abstract
| Publisher Full Text
- 19.
Grunstein M:
Histone acetylation in chromatin structure and transcription.
Nature.
1997; 389(6649): 349–52. PubMed Abstract
| Publisher Full Text
- 20.
Gilmour PS, Rahman I, Donaldson K, et al.:
Histone acetylation regulates epithelial IL-8 release mediated by oxidative stress from environmental particles.
Am J Physiol Lung Cell Mol Physiol.
2003; 284(3): L533–40. PubMed Abstract
| Publisher Full Text
- 21.
Schmeck B, Beermann W, van Laak V, et al.:
Intracellular bacteria differentially regulated endothelial cytokine release by MAPK-dependent histone modification.
J Immunol.
2005; 175(5): 2843–50. PubMed Abstract
- 22.
Goodman RB, Strieter RM, Martin DP, et al.:
Inflammatory cytokines in patients with persistence of the acute respiratory distress syndrome.
Am J Respir Crit Care Med.
1996; 154(3 Pt 1): 602–11. PubMed Abstract
| Publisher Full Text
- 23.
Takizawa H, Ohtoshi T, Kawasaki S, et al.:
Diesel exhaust particles induce NF-kappa B activation in human bronchial epithelial cells in vitro: importance in cytokine transcription.
J Immunol.
1999; 162(8): 4705–11. PubMed Abstract
- 24.
Yamauchi Y, Okazaki H, Desaki M, et al.:
Methotrexate induces interleukin-8 production by human bronchial and alveolar epithelial cells.
Clin Sci (Lond).
2004; 106(6): 619–25. PubMed Abstract
| Publisher Full Text
- 25.
Arae K, Hirata M, Kurata S, et al.:
Mycoplasma pneumoniae induces interleukin-8 production via the epidermal growth factor receptor pathway.
Microbiol Immunol.
2011; 55(10): 748–50. PubMed Abstract
| Publisher Full Text
- 26.
Schleimer RP, Kato A, Kern R, et al.:
Epithelium: at the interface of innate and adaptive immune responses.
J Allergy Clin Immunol.
2007; 120(6): 1279–84. PubMed Abstract
| Publisher Full Text
| Free Full Text
- 27.
Miyata Y, Towatari M, Maeda T, et al.:
Histone acetylation induced by granulocyte colony-stimulating factor in a map kinase-dependent manner.
Biochem Biophys Res Commun.
2001; 283(3): 655–60. PubMed Abstract
| Publisher Full Text
- 28.
Sullivan JT, Richards CS, Lloyd HA, et al.:
Anacardic acid: molluscicide in cashew nut shell liquid.
Planta Med.
1982; 44(3): 175–7. PubMed Abstract
| Publisher Full Text
- 29.
Schmeck B, Lorenz J, N'quessan PD, et al.:
Histone acetylation and flagellin are essential for Legionella pneumophila-induced cytokine expression.
J Immunol.
2008; 181(2): 940–7. PubMed Abstract
- 30.
Hemshekhar M, Sebastin Santhosh M, Kemparaju K, et al.:
Emerging Roles of Anacardic Acid and Its Derivatives: A Pharmacological Overview.
Basic Clin Pharmacol Toxicol.
2011. PubMed Abstract
| Publisher Full Text
- 31.
Asadi L, Eurich DT, Gamble JM, et al.:
Guideline adherence and macrolides reduced mortality in outpatients with pneumonia.
Respir Med.
2012; 106(3): 451–8. PubMed Abstract
| Publisher Full Text
- 32.
Mouktaroudi M, Giamarellos-Bourboulis EJ:
Macrolides for the therapy of nosocomial infections.
Curr Opin Infect Dis.
2012; 25(2): 205–10. PubMed Abstract
| Publisher Full Text







Comments on this article Comments (1)