Keywords
gene cassette, clase 1 integrons, fused gene cassette, transcription
gene cassette, clase 1 integrons, fused gene cassette, transcription
Taking into account the Dr. Partridge recommendations, a new version of figure 2 was provided in order to make clearer the differences between the canonical and truncated GES-1 attC sites. In this new version of the figure, all points raised by Dr. Partridge were addressed.
See the authors' detailed response to the review by Béatrice Berçot
See the authors' detailed response to the review by Sally Partridge
Class 1 integrons are capable of inserting, excising and rearranging gene cassettes by a site-specific recombination mechanism. These assembly platforms can also act as expression systems due to the presence of a promoter region (Pc), which drives the expression of genes captured by integron1. Moreover, naturally occurring integrons may have a second promoter (P2), which is activated by the insertion of three G residues between -35 and -10 hexamers1. Gene cassettes are generally promoterless units associated with a recombination site (attC or 59-be), which confers the ability of each structure to be mobilized independently2, and the Left Hand (LH – 1L and 2L core sites) and Right Hand (RH – 1R and 2R core sites) domains from attC sites are crucial for this mobilization3. Studies focusing on gene cassette translation in the context of integrons are rare, however, it was recently showed that attC sites regulate the translation of downstream cassettes due to their peculiar sequences composed by imperfect inverted repeats that form stem-loop structures. These secondary structures prevent ribosome progression throughout mRNA, reflecting in a decreased expression of more distal genes regarding Pc4. Conversely, TIR (Translation Initiation Region)-deficient gene cassettes could have their expression promoted by the presence of ORF-115. This ORF, when present, is found at the attI site preceding the gene cassette array. It codes for 11 amino acids and harbours its own Shine-Dalgarno (SD) sequence. Therefore, the ORF-11 recruits the ribosomes and, through an event of coupled translation, the subsequent TIR-deficient gene cassette could be expressed4,5. Gene cassettes can be found inserted in integrons or in other secondary sites, or free in the cytoplasm as a closed circle, in which the 5’ end (5’ UTR) and the attC recombination site are covalently linked6.
As demonstrated previously, several stress conditions could evoke the activation of the SOS response resulting in integron-integrase expression7. Therefore, under stress, the integrase activity increases, favoring the occurrence of integration/excision/rearrangements events.
Although rare, fused cassettes may be generated by partial or total loss of the first attC, retaining both complete coding regions and, therefore, creating permanent gene arrays comparable to bacterial operons8. The functionality of such structures has been indirectly inferred by the resistance profile of transformants carrying the fusion9; however, the transcription itself has never been verified.
This study showed the dynamics of fused cassette mobilization, the co-transcription of the gcu14-blaGES-1/aacA4 cassette array and the effect of cassette position on transcription levels in Pseudomonas aeruginosa wild lineages carrying class 1 integrons. Moreover, the presence of translation signals in this gene cassette array was determined.
An unknown Open Reading Frame (ORF), gcu14 (gene cassette of unknown function), followed by the fused cassette blaGES-1/aacA4, created by partial loss of GES-1 attC were present in integrons from clinical P. aeruginosa isolates (PS1 and PS26)10. Total RNA was extracted and purified according to the manufacturer’s instructions with the SV 96 Total RNA Isolation System (Promega). Northern blot using 7 μg of total RNA from PS1 and PS26 was performed in order to detect the transcript originated from gcu14-blaGES-1/aacA4 cassette array. After electrophoresis in a denaturing-formaldehyde 1.5% agarose gel, the total RNA was transferred to the Hybond-N+ nylon membrane (GE Healthcare) by upward capillary transfer. An amplicon of 519bp corresponding to part of the blaGES-1 gene was used as a probe (Table 1) in hybridization assay. The GES probe was labeled with the AlkPhos Direct Labelling kit (GE Healthcare) and hybridized with the target RNA immobilized on the Hybond-N+ membrane as recommended. The chemiluminescence was detected with the CDP-Star detection reagent (GE Healthcare) according to manufactures. Immediately after applying the detection reagents, the blot was drained, incubated five minutes at room temperature and exposed to the Hyperfilm ECL (GE Healthcare) for 60 minutes at room temperature.
In order to verify whether the relative position of gene cassettes on the variable region plays a role in transcription level, real-time RT-PCR reactions using the TaqMan System (Applied Biosystems) were performed with primers and probes detailed in Table 1. The rpsL gene of the P. aeruginosa chromosome was amplified by PCR (Table 1) and used as a reference gene for normalization. The relative quantification (RQ) results were presented as ratios of gene transcription between the target gene (cassettes) and the reference gene (rpsL), which were obtained according to the following equation: RQ=2-ΔCT, where CT is the value corresponding to the crossing point of the amplification curve with the threshold and ΔCT=CT target gene minus CT reference gene. The effect of cassette position on gene transcription was considered significant when the ratios obtained between RQ values (RQ value of cassette 1/RQ value of cassette 2) were ≥2.0, taking into account the standard deviation intervals.
In order to induce cassette excision from integrons, PS1 and PS26 strains10 were submitted to thermal stress during the log growth phase to induce integrase activity. Cells were grown on Luria-Bertani (LB) broth medium (OXOID) at 37°C for two hours. Subsequently, the bacterial cultures were submitted to a heat shock at 4°C for 30 minutes and immediately incubated at 42°C for another 30 minutes. Briefly, the total DNA from PS1 and PS26 cultured under thermal stress were obtained with the Wizard Genomic DNA purification kit (Promega) following manufacturer recommendations and used as templates in inverse PCR reactions. The inverse PCR was performed with primers facing outwards towards the ends of gcu14, blaGES-1 and aacA4 so that only circular gene cassette configurations would be amplified. The reactions targeting the circular forms of gcu14, blaGES-1, aacA4 and blaGES-1/aacA4 fusion was performed with primers and combinations described in Table 1. The inverse PCR was performed using Platinum Taq DNA Polymerase reagents (Invitrogen), and the following components were added to a sterile 0.2-mL tube: 5 µL of 10X PCR buffer (1X final concentration); 1 µL of 10mM dNTP mixture (0.2 mM each); 1.5 µL of 50mM MgCl2 (1.5 mM final concentration); 2 µL of 15 µM of each primer (30 µM each); 100 ng of template DNA; 0.3 µL of Platinum Taq DNA Polymerase (1U final concentration). The tubes were incubated in the Eppendorf MasterCycler (Eppendorf) at 94°C for 2 minutes and PCR amplification was performed in 40 cycles consisting of: 94°C for 30 seconds; 55°C for 30 seconds; and 72°C for 3 minutes. The amplicons generated with the inverse PCR were purified using Wizard SV Gel and PCR Clean-Up system kit (Promega) and directly sequenced on both strands. Sequencing reactions were performed with Big Dye Terminator RR Mix (Applied Biosystems) in an ABI 3730 XL DNA Analyzer (Applied Biosystems). Nucleotide sequences were compared to those available in the GenBank database accessible on the National Center for Biotechnology Information website (http://www.ncbi.nlm.nih.gov). All primers used in PCR, sequencing and real time RT-PCR are described in Table 1.
Analyses in silico were performed to search for a potential promoter for gcu14 gene cassette. The 5’UTR from gcu14 were submitted to the promoter predictor programs Neural Network for Promoter Prediction version 2.2 (Berkeley Drosophila Genome Project, http://www.fruitfly.org/index.html) and BPROM (SoftBerry, http://linux1.softberry.com/berry.phtml). Results with the highest scores were selected as candidates for a putative promoter.
The integrons analysed in this study harbored the weak Pc configuration (PcW)11, and they carry the gcu14 as the first gene cassette (see reference 12 for nomenclature), which has not been reported so far. Considering that transcription initiates from the Pc promoter placed upstream the cassette array, both monocistronic and full length polycistronic transcripts could be identified. In fact, Northern blot and hybridization assays revealed a unique signal of approximately 2,300 bases, which corresponds to the co-transcription of the entire array (gcu14-blaGES-1/aacA4) (Figure 1). This result is in agreement with previous work in which the occurrence of transcripts containing more than one gene cassette was observed by Northern blot analysis1. Moreover, this finding gives support to the lack of attC function in terminating transcription of downstream gene cassettes as demonstrated previously4.

The full length transcript (2,300 bases), corresponding to the entire gene cassette array, hybridized with the GES probe (arrow). The fragment sizes of the RNA marker (Promega) used in RNA electrophoresis are indicated.
This fusion retained both entire coding regions and, due to a possible erroneous recombination event, the blaGES-1 attC site was replaced by part of the attI1 site (DQ236170)10, reducing it to the 6bp from the 1L core site (Figure 2). Taking into account that the region responsible for stem-loop formation was missing in GES-1 attC and the participation of this site in terminating translation4, our findings indirectly suggested that blaGES-1 and aacA4 translation is occurring in a unique step.

The underlined uppercase letters represent the blaGES-1 and aacA4 partial coding regions. Italicized lowercase letters highlighted in grey define the attC sites, and their core sites (1L, 2L, 2R, 1R) are in boldface and underlined. The vertical arrows show the recombination point and the beginning of the next cassette.
The strains were submitted to thermal stress conditions in order to verify the dynamics of mobilization of gcu14-blaGES-1/aacA4 gene cassettes. Since the excision event depends on the recognition of the LH and RH domains of the attC site, and that the 2L core site and the entire RH domain are missing in GES-1 attC (Figure 2), it is expected that the blaGES-1/aacA4 excision occurs only at the aacA4 attC site, and that this structure is excised together as a unique cassette.
Positive results were obtained for the gcu14, gcu14-aacA4 and gcu14-blaGES-1/aacA4 circular forms, but not for blaGES-1 and aacA4 alone. This finding indicates that the GES-1 attC is not functional and that the fused gene cassette is excised as a unique cassette. Moreover, the presence of gcu14-aacA4 circular form suggests that this strain carries a second integron containing this gene cassette arrangement. Sequencing assessed the recombination point where excision occurred, confirming the occurrence of free circular forms (Figure 3). This is in agreement with the presence of the PcW configuration, in which the corresponding intI1 gene codes for a high efficient integrase11. The lack of activity of a truncated attC had also been observed before when associated with aadA1013. However, Ramirez and colleagues14 showed that the integrase was able to recognize and mediate excision of a truncated site associated to aadA1, indicating that the genetic context of such truncated sites could influence their role in IntI1 recognition and mobilization.
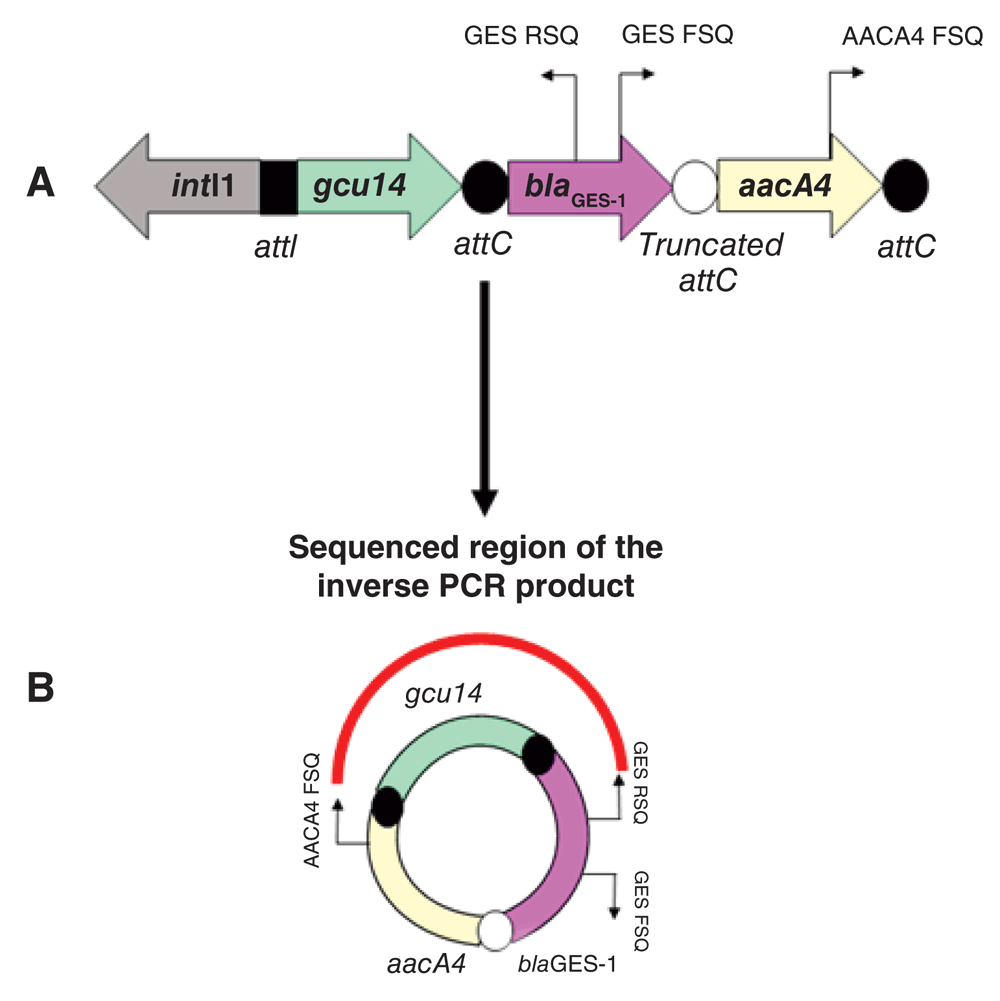
(A) inserted/linearized form of integron harbouring the fused cassettes. Arrows show the gene transcription orientation. Thin arrows represent the annealing sites of the inverse PCR and sequencing primers whose generated product corresponded to the cassette circular form illustrated in (B). (B) Illustration of the free circular form of gene cassettes represented in (A). Thin arrows show the primers used to obtain the inverse PCR product (GES FSQ and GES RSQ) and the primers used in sequencing (AACA4 FSQ and GES RSQ) that revealed the excision in block of the entire gene cassette arrangement (gcu14-blaGES-1/aacA4) from integron (red curved line).
The relative quantification performed by real time RT-PCR revealed that PS1 and PS26 presented very similar RQ values for gcu14-blaGES-1/aacA4 transcription (Figure 4). This result was expected since integrons from these two strains have the same backbone, including the Pc promoter, and are at the same genetic environment10.

The relative quantification values obtained by real time RT-PCR are indicated for each gene cassette and for the fusion blaGES-1/aacA4 when considered as a unique gene.
gcu14, the first cassette in integrons with the weak PcW configuration, presented approximately two-fold higher transcription when compared to blaGES-1 and aacA4 separately or when the fused cassette blaGES-1/aacA4 was considered (Figure 4). The same RQ value obtained for blaGES-1, aacA4 and the fusion reveals that these two ORFs are transcribed as a unique gene. The lower transcript amount of blaGES-1/aacA4 compared to gcu14 lies on the distance between these gene cassettes and Pc, which is one of the determinants influencing cassette transcription1,7, and it shows the effect of cassette position on expression levels.
A putative promoter for gcu14 (-35 TTGATG [17 bp] -10 TGTTAC) was found 45 bp upstream from its start codon, which has the potential to influence transcription. Moreover, the ORF-11, which enhances the translation efficiency of downstream TIR-deficient cassettes inserted in integrons5, was found at the attI1 region preceding the TIR-deficient gcu14 gene cassette. This ORF contained its own Shine-Dalgarno (SD) sequence placed 8 bp upstream of the ATG codon. The ribosome at the ORF-11 stop codon could, therefore, be carried along the mRNA by lateral diffusion, reinitiating translation at the gcu14 start codon. A potential SD sequence was identified 10 bp upstream of the fused cassette blaGES-1/aacA4. In addition, the loss of the GES-1 attC region, which is involved in stem-loop formation, may enhance the chances of aacA4 translation, since this attC, reduced to the 6bp of the 1L core site, no longer constitutes a physical barrier to ribosome progression4. Together, these findings create a scenario for the occurrence of gcu14-blaGES-1/aacA4 expression in PS1 and PS26, which then provides a possible explanation for their resistance profile to β-lactams and aminoglycosides that has been observed elsewhere10.
Fused cassettes have been found in class 1 integrons9,13–18; however, the transcription of such structures has rarely been addressed. This work showed the transcription pattern of a fused cassette as a polycistronic mRNA and that these unusual structures are excised as a unique cassette. The mobilization in block of the entire gcu14-blaGES-1/aacA4 array together with its active transcription, and the presence of translational signatures demonstrate the potential for dissemination and expression of multidrug resistance, in a one-step fashion, to other bacteria. Therefore, such events could represent a threat to public health and to the establishment of efficient antibiotic regiments.
The sequence of the cassette array composed by the fusion has been deposited in the GenBank database under accession number DQ236170. The sequence obtained from the inverse PCR amplicon, showing the circular form gene organization, was submitted to GenBank under accession number KT336477.
Figshare: Polycistronic transcription of fused cassettes and identification of translation initiation signals in an unusual gene cassette array from Pseudomonas aeruginosa doi: http://dx.doi.org/10.6084/m9.figshare.65171719
ELF and ACPV conceived the study and designed the experiments. ELF carried out the research and prepared the first draft of the manuscript. All authors were involved in the revision of the draft manuscript and have agreed to the final content.
This work was supported by the CNPq and FAPERJ fellowship and Oswaldo Cruz Institute Grant.
We thank those involved in PDTIS platform to enable us to do the sequencing and real-time relative quantification analysis work.
| Views | Downloads | |
|---|---|---|
| F1000Research | - | - |
|
PubMed Central
Data from PMC are received and updated monthly.
|
- | - |
Competing Interests: No competing interests were disclosed.
Competing Interests: No competing interests were disclosed.
Competing Interests: No competing interests were disclosed.
Competing Interests: No competing interests were disclosed.
Competing Interests: No competing interests were disclosed.
Alongside their report, reviewers assign a status to the article:
| Invited Reviewers | ||
|---|---|---|
| 1 | 2 | |
|
Version 3 (revision) 27 Nov 15 |
read | |
|
Version 2 (revision) 07 Sep 15 |
read | read |
|
Version 1 28 Mar 13 |
read | read |
Provide sufficient details of any financial or non-financial competing interests to enable users to assess whether your comments might lead a reasonable person to question your impartiality. Consider the following examples, but note that this is not an exhaustive list:
Sign up for content alerts and receive a weekly or monthly email with all newly published articles
Already registered? Sign in
The email address should be the one you originally registered with F1000.
You registered with F1000 via Google, so we cannot reset your password.
To sign in, please click here.
If you still need help with your Google account password, please click here.
You registered with F1000 via Facebook, so we cannot reset your password.
To sign in, please click here.
If you still need help with your Facebook account password, please click here.
If your email address is registered with us, we will email you instructions to reset your password.
If you think you should have received this email but it has not arrived, please check your spam filters and/or contact for further assistance.
Besides the comments already made by the reviewers, I would like to specify some points:
- The C to G mutations that converts the usual weak PcW variant into PcWTGN-10 has
... Continue reading Dear Authors,Besides the comments already made by the reviewers, I would like to specify some points:
- The C to G mutations that converts the usual weak PcW variant into PcWTGN-10 has been shown to increase (instead of 'decrease', first part of the Results section).
Response: This sentence was removed.- I have not checked which IntI1 had been used for the excision assay, usually these are made with the most efficient "PcW" IntI1 (IntI1R32_H39); note that in our PLoS Genetics paper (Jove et al. 2010) we evidenced the IntI1 "PcWTGN-10" (IntIP32_H39) were much less efficient for excision of gene cassettes, which could had been taken into account in your discussion.
Response: As properly noticed by you, our Pc configuration is the weak one, and this was changed in the text. However, we agree that including this discussion we will improve our manuscript (lines 182 and 184).- The hypothesis of a synergistic effect between the Pc and P(gcu14) tandem promoters does not fit the observed phenotypes: since the resulting level of transcription observed for the downstream genes is lower I suggest that some transcriptional interferences occur rather than synergistic interactions. By the way in absence of any experimental evidence for the functionality of the gcu14 promoter, such hypothesis is overstated. It would have been interesting to check the level of transcription of the blaGES-1,aacA4 gene cassette when gcu14 is deleted.
Response: we totally agree with this observation, in fact our conclusion is overstated since no experimental assay was performed to test the promoter synergy. Therefore, we remove this part from the text. We also agree that would be interesting to verify the expression of the fused cassette in the absence of gcu14, however it was not the focus of this paper.- First, the class 1 integron sequence of the PS1 strain in Genbank DQ236170 does not display the rare C to G mutation that converts a weak "PcW" promoter into a stronger "PcWTGN-10" as stated in the paper. Consequently you should write "the weak PcW configuration" instead of "weak PcWTGN-10". Also it means that its own IntI1 integrase is likely to be optimally efficient.
Response: Dr. Jové is absolutely right, the DQ236170 accession number presented the PcW configuration and not the PcWTGN-10 as stated in the manuscript, we apologize about this mistake. This was corrected in the text.
- Then, the sequence of the class 1 integron as deposited in the Genbank does not display the complete array of gene cassette (the end of the attCaacA4 is not available) which means that it is ambigous whether there is a downstream third gene cassette or not. Consistently this array of gene cassette, although a novel one, has not been numbered in the INTEGRALL database (http://integrall.bio.ua.pt/?acc=DQ236170)
Response: We know that aacA4 was the last cassette in the array because this variable region was obtained using primers annealing in the attI1 and in the beginning of qacE∆1 from 3’CS. It is true that the aacA4 attC is not completed in the GenBank accession number DQ236170. Therefore, spite of the incomplete attC sequence we are sure that the aacA4 is the last cassette from this array.
- Lastly, as an element of reply to the reviewer Dr Bercot, I would like to notify that the so-called gcu14 GC has not been reported in other reports except in an environmental strain of Citrobacter (and the sequence is not 100% identical, Genbank FM998050).
Response: thanks for this observation. It was included in the text.
Besides the comments already made by the reviewers, I would like to specify some points:
- The C to G mutations that converts the usual weak PcW variant into PcWTGN-10 has been shown to increase (instead of 'decrease', first part of the Results section).
Response: This sentence was removed.- I have not checked which IntI1 had been used for the excision assay, usually these are made with the most efficient "PcW" IntI1 (IntI1R32_H39); note that in our PLoS Genetics paper (Jove et al. 2010) we evidenced the IntI1 "PcWTGN-10" (IntIP32_H39) were much less efficient for excision of gene cassettes, which could had been taken into account in your discussion.
Response: As properly noticed by you, our Pc configuration is the weak one, and this was changed in the text. However, we agree that including this discussion we will improve our manuscript (lines 182 and 184).- The hypothesis of a synergistic effect between the Pc and P(gcu14) tandem promoters does not fit the observed phenotypes: since the resulting level of transcription observed for the downstream genes is lower I suggest that some transcriptional interferences occur rather than synergistic interactions. By the way in absence of any experimental evidence for the functionality of the gcu14 promoter, such hypothesis is overstated. It would have been interesting to check the level of transcription of the blaGES-1,aacA4 gene cassette when gcu14 is deleted.
Response: we totally agree with this observation, in fact our conclusion is overstated since no experimental assay was performed to test the promoter synergy. Therefore, we remove this part from the text. We also agree that would be interesting to verify the expression of the fused cassette in the absence of gcu14, however it was not the focus of this paper.- First, the class 1 integron sequence of the PS1 strain in Genbank DQ236170 does not display the rare C to G mutation that converts a weak "PcW" promoter into a stronger "PcWTGN-10" as stated in the paper. Consequently you should write "the weak PcW configuration" instead of "weak PcWTGN-10". Also it means that its own IntI1 integrase is likely to be optimally efficient.
Response: Dr. Jové is absolutely right, the DQ236170 accession number presented the PcW configuration and not the PcWTGN-10 as stated in the manuscript, we apologize about this mistake. This was corrected in the text.
- Then, the sequence of the class 1 integron as deposited in the Genbank does not display the complete array of gene cassette (the end of the attCaacA4 is not available) which means that it is ambigous whether there is a downstream third gene cassette or not. Consistently this array of gene cassette, although a novel one, has not been numbered in the INTEGRALL database (http://integrall.bio.ua.pt/?acc=DQ236170)
Response: We know that aacA4 was the last cassette in the array because this variable region was obtained using primers annealing in the attI1 and in the beginning of qacE∆1 from 3’CS. It is true that the aacA4 attC is not completed in the GenBank accession number DQ236170. Therefore, spite of the incomplete attC sequence we are sure that the aacA4 is the last cassette from this array.
- Lastly, as an element of reply to the reviewer Dr Bercot, I would like to notify that the so-called gcu14 GC has not been reported in other reports except in an environmental strain of Citrobacter (and the sequence is not 100% identical, Genbank FM998050).
Response: thanks for this observation. It was included in the text.